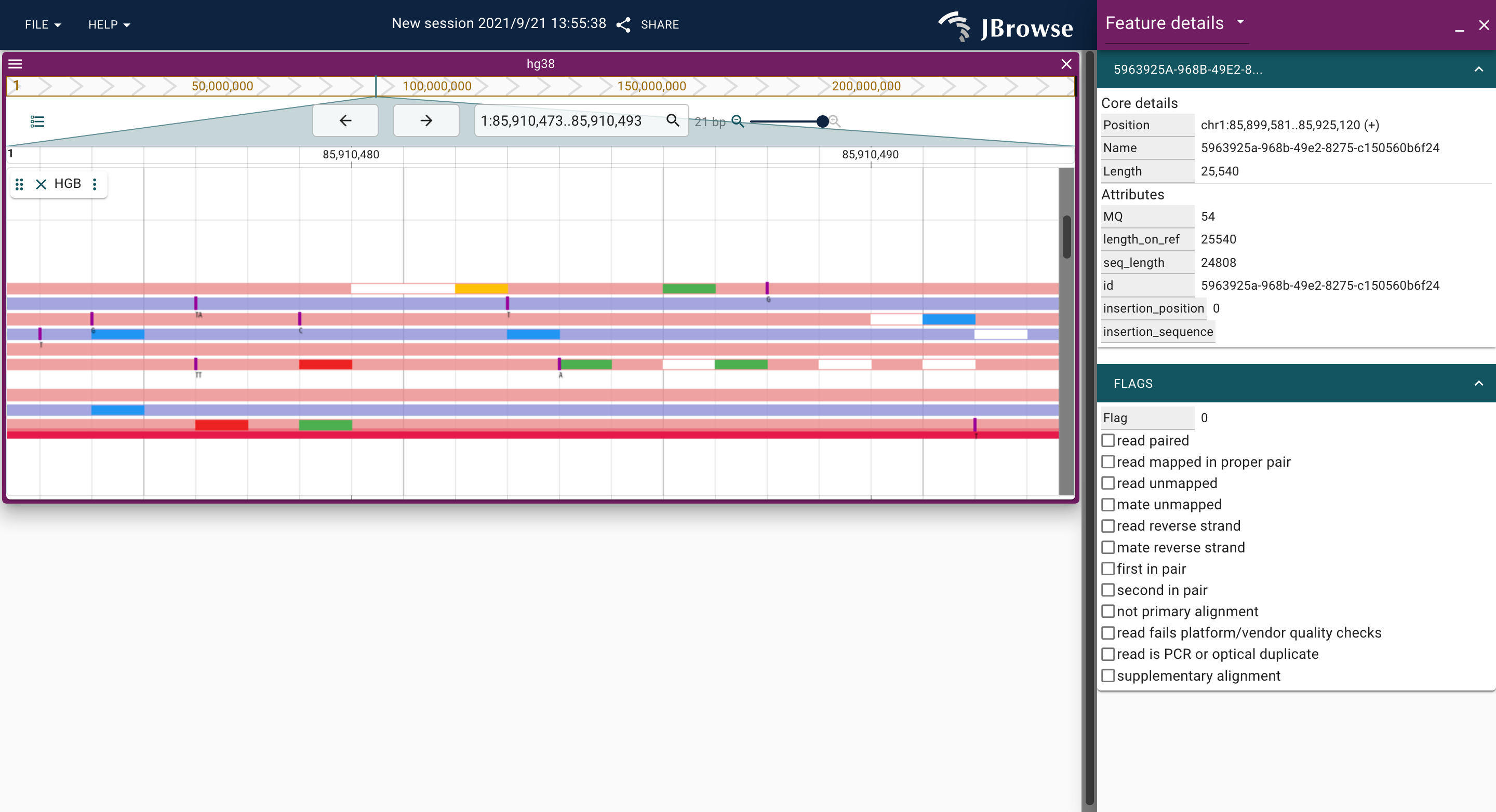
JBrowse2 plugin for displaying long-read alignments by hgb.
- Rust nightly (cargo)
- Node.js
The input bam file needs to be attached MD tags by samtools calmd and indexed by samtools index.
- Docker
docker run --rm -ti -p 9000:9000 -v `pwd`:/data 6thbridge/jbrowse2-hgb:master /data/<input.bam>- Singularity
singularity build jb2-hgb.sif docker://6thbridge/jbrowse2-hgb:master
singularity -s run jb2-hgb.sif <input.bam>Access to http://localhost:9000/static/index.html.
bash -x start_hgb_server.sh $NUM_OF_THREADS $LOCATION_OF_BAM
bash -x start_hgb_server.sh $NUM_OF_THREADS $LOCATION_OF_BAM 0.0.0.0:5000The following command needs to run on another shell.
bash -x start_jbrowse_hgb.sh
bash -x start_jbrowse_hgb.sh $HGB_SERVER_URL 38
bash -x start_jbrowse_hgb.sh $HGB_SERVER_URL 19You can specify reference genome as the second argument as 38 or 19 (hg38 or hg19, respectively).
Add to the "plugins" of your JBrowse Web config. The unpkg CDN should be stable, or you can download the js file to your server
{
"plugins": [
{
"name": "hgb",
"url": "https://unpkg.com/jbrowse-plugin-hgb/dist/jbrowse-plugin-hgb.umd.production.min.js"
}
]
}This plugin is currently quite basic, and there is no mouseover interactivity or drawn labels on features
See DEVELOPMENT