neXtProt - The knowledge resource on human proteins
This is a code repository for the SIB - Swiss Institute of Bioinformatics CALIPHO group neXtProt project
neXtProt feature viewer
The feature viewer is a super easy javascript library to use in order to draw the different features covering a sequence for a better visualization.
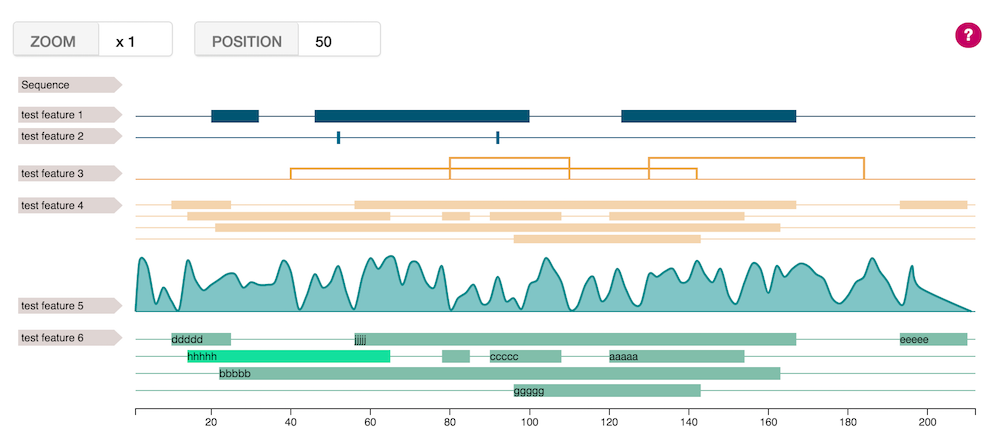
Live demo : https://cdn.rawgit.com/calipho-sib/feature-viewer/v1.0.0/examples/index.html
Getting Started
1. You can get the library in your project using bower or npm
//BOWER//
bower install feature-viewer
//NODE//
npm install feature-viewer
Or Include the feature-viewer JS and CSS from rawgit CDN in the header of your html
NOTE : If you already got the dependencies (D3, Bootstrap & Jquery) in your project, use the simple minified version instead of the bundle :
2. Specify a div in your html
<div id="fv1"></div>
3. Create an instance of FeatureViewer in javascript with the sequence (or a length), the div in which it will be display and the rendering options of your choice.
//For Node add before : var FeatureViewer = require("feature-viewer"); var ft = 'MALWMRLLPLLALLALWGPGPGAGSLQPLALEGSLQKRGIVEQCCTSICSLYQLE' '#fv1' showAxis: true showSequence: true brushActive: true //zoom toolbar:true //current zoom & mouse position bubbleHelp:true zoomMax:50 //define the maximum range of the zoom ; //Instead of a sequence, you can also initialize the feature viewer with a length (integer) :var ft = 213'#fv1';4. Finally, add the features
ft;5. Et voila!
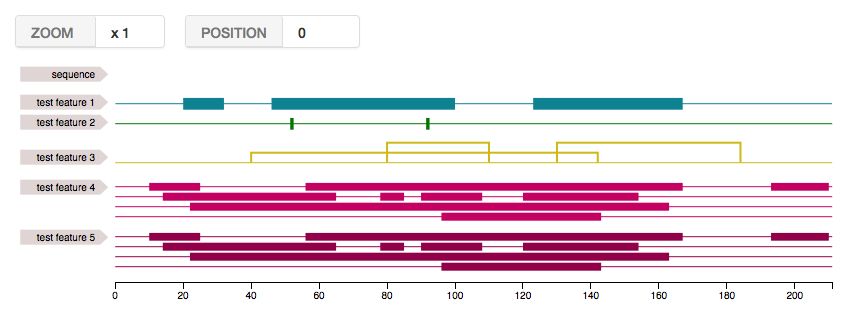
Functionalities
-
Zoom into the Feature-viewer by selecting a part of the sequence with your mouse. Zoom out with a right-click.
You can also zoom programmatically with the methodszoom(start,end)andresetZoom() -
A tooltip appears when the mouse is over a feature, giving its exact positions, and optionally, a description.
-
beside the positions for each element, you can also give a description & an ID, allowing you to link click event on the feature to the rest of your project.
Options
- Show axis
- Show sequence
- Brush active (zoom)
- Toolbar (current zoom & position)
- Bubble help
- Zoom max
- Features height
- Offset
ClearInstance()
You may sometimes want to reload your feature-viewer with new parameters. To avoid memory leaks, the method clearInstance() will clear each element & listener for you before you delete the feature-viewer instance.
Documentation
Check out this page for a better understanding of how to use the feature viewer and its possibilities :
Use it with NeXtProt API
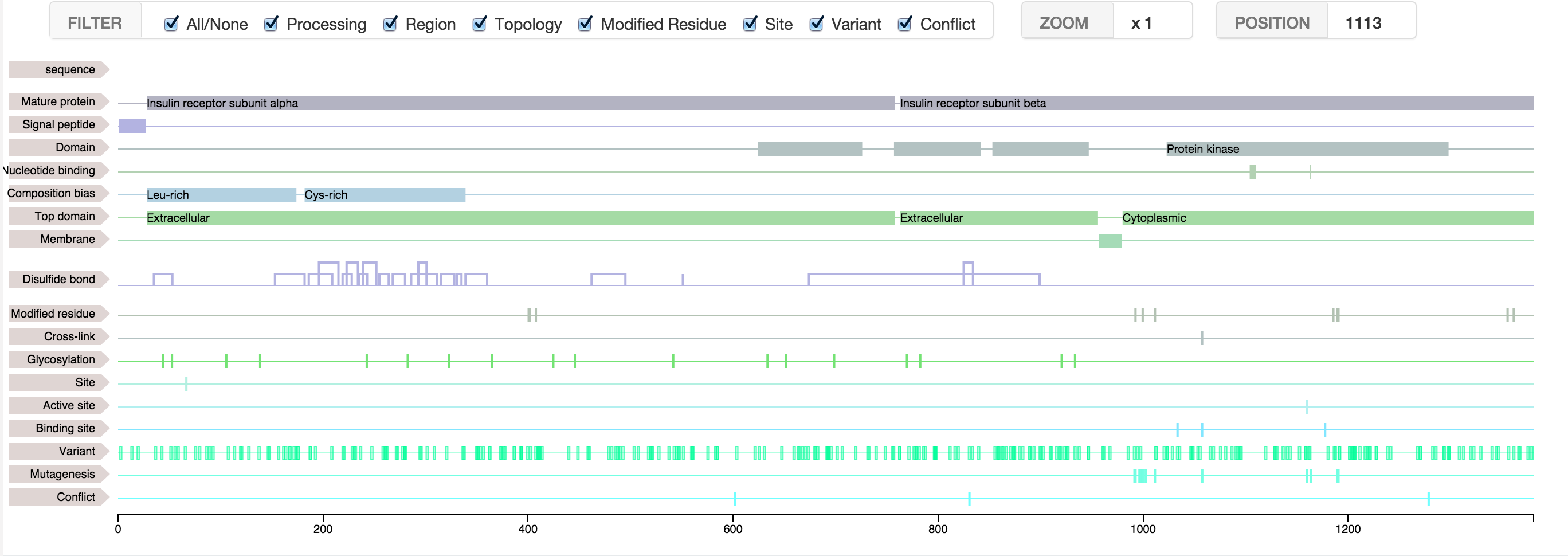
It is possible to fill the feature viewer with protein features from NeXtProt, the human protein database.
- First, find your protein of interest in NeXtProt and get the neXtProt accession (NX_...). (You can find your protein by entering an accession number of another database, like UniProt or Ensembl)
- Then, check the type of feature in the NeXtProt API that you would like to add to your viewer. For example, "propeptide" or "mature-protein".
- Include the feature viewer bundle with nextprot to your html : feature-viewer.nextprot.js
- Finally, create your feature-viewer like this :
//initalize nextprot Clientvar applicationName = 'demo app'; //please provide a name for your applicationvar clientInfo='calipho group at sib'; //please provide some information about youvar nx = applicationName clientInfo; //var entry = "NX_P01308";var isoform = "NX_P01308-1"; // feature viewer optionsvar options = showAxis: true showSequence: truebrushActive: true toolbar:truebubbleHelp: true zoomMax:20 ; // Create nextprot feature viewerExamples
https://search.nextprot.org/entry/NX_P01308/view/proteomics
Support
If you have any problem or suggestion please open an issue here.
Development
git clone https://github.com/calipho-sib/feature-viewer.git
npm install (will install the development dependencies)
bower install (will install the browser dependencies)
...make your changes and modifications...
npm run dist (will create the min & bundle versions in dist/)
npm run build (will create the bundle js & css in build/ for node)
grunt bump (will push and add a new release)
npm publish (will publish in npm)
License
Copyright (c) 2015, SIB Swiss Institute of Bioinformatics
This program is free software; you can redistribute it and/or modify it under the terms of the GNU General Public License as published by the Free Software Foundation; either version 2 of the License, or (at your option) any later version.