d3-electrophoresis
A D3.js plugin to simulate electrophoresis which is a basic biotechnological technique to visualize length of DNA segments cut by restriction enzyme. Using this plugin, you can cut not only DNA sequence, but also any text, by words.
Examples
Cutting text by words
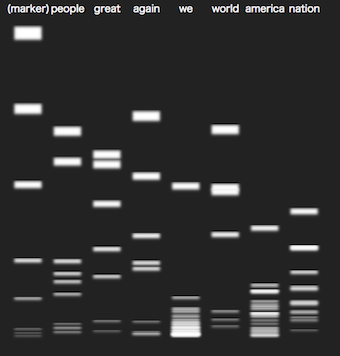
You can change text and words in the showcases.
- Simple example: Showcase and code.
- Changinge scale: showcase and code.
- Trump inauguration text: Showcase, Code.
- DNA sequence: Showcas, Code.
Using SVG for CGI
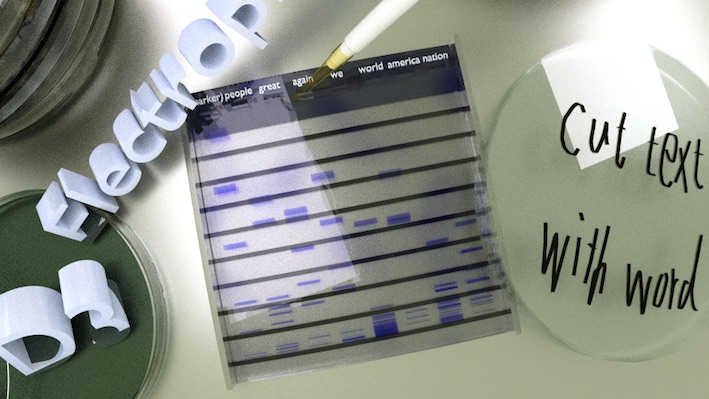
By SVG Crowbar, you can download the resulted DNA bands as SVG. And the SVG can be used for, such as 3D object in Blender. The CGI image below was made with Blender, including SVG bands in the gel.
The detail and other images are on the blog: New D3.js plugin: D3-electrophoresis released.
Install
- Download d3-electrophoresis.js from right click, or use npm
npm install d3-electrophoresisand get it in "src" folder. - In HTML, load D3.js (v.4) and d3-electrophoresis.js with
<script>tags, like the code below.
<script src="https://cdnjs.cloudflare.com/ajax/libs/d3/4.5.0/d3.min.js" charset="utf-8"></script>
<script src="d3-electrophoresis.js"></script>
Basic Usage
var gel = ; var svg = d3 ; // Make black background first. var text = "AGCGCCCAATACGCAAACCGCCTCTCCCCGCGCGTTGGCCGATTCACGTTTACGAGTTGGAAACGA";var words = 1510152030405070100"CAAA""GTTGG""GGATCC" "GA""GT""CCT""ACC" "GAAA";var names = "marker" "1" "2" "3" "4" "5" "6" "7";// You can set every parameter by dictionary or method-chain. Example: {DNA: text} or .DNA(text).gel = ;svg;Parameters
You can set every parameter by dictionary input:
var gel = ;and/or by method-chain:
var gel = ;Tips
Changing scale

To change vertical scale of band, like changing gel concentration, use .scale() chain, and input d3's scale object.
For example, to make band within y=200~400 in linear scale, add scale() chain after electrophoresis().~, and input d3-scale object:
var yScale = d3;gel = ;API Reference
# electrophoresis.DNA([text])
Target text to be cut.
Default: "AGCGCCCAATACGCAAACCGCCTCTCCCCGCGCGTTGGCCGATTCACGTTTACGAGTTGGAAACGA".
# electrophoresis.enzymes([list])
Word list to cut target text.
When the word list includes a number list as a element, (e.g. ["AC",[10,15,20]]), the numbers are directly used as resulted length of cut segmented text without cutting text. When the word list includes a word list as a element, (e.g. ["AC",["GT","CCT"]]), target text is cut by both of the two words all together.
Default: [[1,5,10,15,20,30,40,50,70,100],"CAAA","GTTGG","GGATCC", "GA",["GT","CCT"],"ACC", "GAAA"].
# electrophoresis.names([list])
Name list of enzymes. They appear as tooltips on each resulted lane.
Default: ["marker", "1", "2", "3", "4", "5", "6", "7"].
# electrophoresis.scale([obj])
A scale to map length values of segmented texts from top to the bottom. The input must be a scale object in D3-v.4 syntax, such as d3.scaleSqrt(), d3.scaleLinear(), d3.scalePow() .etc. See how to use d3/d3-scale.
To make band within height of 200-500 in linear scale, for example, use d3.scaleLinear().range([200, 500])).
Default: d3.scaleLinear().
# electrophoresis.lane_number([int])
Max number of lanes.
Default: 8.
# electrophoresis.gel_margin([dict])
Margin size from edges of background.
Default: {top: 40, right: 20, bottom: 30, left: 20}.
# electrophoresis.duration([float])
Time to finish animation.
Default: 6000.
# electrophoresis.band_width([float])
Width of bands.
Default: 38.
# electrophoresis.band_blur([float])
Blur of bands.
Default: 2.
# electrophoresis.band_thick_min([float])
Minimum thickness of bands.
Default: 1.
# electrophoresis.band_thick_rate([float])
How bands turns less thick as the bands go down.
Default: 0.04
# electrophoresis.tooltip_name_on([str])
"on" or "off" for tooltips of enzyme name on upper side.
Default: "on".
# electrophoresis.tooltip_name_size([float])
Size of name tooltips.
Default: 17.
# electrophoresis.tooltip_name_offsetX([float])
Horizontal offset of name tooltips.
Default: 0.
# electrophoresis.tooltip_name_offsetY([float])
Vertical Offset of name tooltips.
Default: 20.
# electrophoresis.tooltip_band_on([str])
"on" or "off" for tooltips of band length.
Default: "on";
# electrophoresis.tooltip_band_size([float])
size of band tooltips.
Default: 13.
# electrophoresis.tooltip_band_offsetX([float])
Horizontal offset of band tooltips.
Default: 0.
# electrophoresis.tooltip_band_offsetY([float])
Vertical offset of band tooltips.
Default: 0.